Inference for Discrete Data
Linking Probability and Data
Discrete Inference
Inference for Binomial Type Variables
How do we learn from data?

It is probably better posed as, what can we learn from data?
The Big Idea
Inference
In the most basic form, learning something about data that we do not have from data that we do. In the jargon of statistics, we characterize the probability distribution of some feature of the population of interest from a sample drawn randomly from that population.
Quantities of Interest:
Single Proportion: The probability of some qualitative outcome with appropriate uncertainty.1
Single Mean: The average of some quantitative outcome with appropriate uncertainty.1
The Big Idea
Inference
In the most basic form, learning something about data that we do not have from data that we do. In the jargon of statistics, we characterize the probability distribution of some feature of the population of interest from a sample drawn randomly from that population.
Quantities of Interest:
Compare Proportions: Compare key probabilities across two groups with appropriate uncertainty.1
Compare Means: Compare averages/means across two groups with appropriate uncertainty.1
The Big Idea
Inference
In the most basic form, learning something about data that we do not have from data that we do. In the jargon of statistics, we characterize the probability distribution of some feature of the population of interest from a sample drawn randomly from that population.
Study Planning
Plan studies of adequate size to assess key probabilities of interest.
Inference for Data
Of two forms:
- Binary/Qualitative
- Quantitative
We will first focus first on the former. But before we do, one new concept.
The standard error
It is the degree of variability of a statistic just as the standard deviation is the variability in data.
The standard error of a proportion is \sqrt{\frac{\pi(1-\pi)}{n}} while the standard deviation of a binomial sample would be \sqrt{n*\pi(1-\pi)}.
The standard deviation of a sample is s while the standard error of the mean is \frac{s}{\sqrt{n}}. The average has less variability than the data and it shrinks as n increases.
Let’s think about an election….
Consider election season. The key to modeling the winner of a presidential election is the electoral college in the United States. In almost all cases, this involves a series of binomial variables.
Almost all states award electors for the statewide winner of the election and DC has electoral votes.
We have polls that give us likely vote percentages at the state level. Call it
p.winfor the probability of winning. We can then deployrbinom(1000, size=1, p.win)to derive a hypothetical election.Rinse and repeat to calculate a hypothetical electoral college winner. But how do we calculate
p-win?We infer it from polling data
Qualitative: The Binomial
Is entirely defined by two parameters, n – the number of subjects – and \pi – the probability of a positive response. The probability of exactly x positive responses is given by:
Pr(X = x | \pi, n) = {n \choose x} \pi^x (1-\pi)^{n-x}
The binomial is the canonical distribution for binary outcomes. Assuming all n subjects are alike and the outcome occurs with \pi probability,1 then we must have a sample from a binomial. A binomial with n=1 is known as a Bernoulli trial
Features of the Binomial
Two key features:
Expectation: n \cdot \pi or number of subjects times probability, \pi \textrm{ if } n=1.
Variance is n \cdot \pi \cdot (1 - \pi) or \pi(1-\pi) \textrm{ if } n=1 and standard deviation is \sqrt{n \cdot \pi \cdot (1 - \pi)} or \sqrt{\pi(1-\pi)} \textrm{ if } n=1.
Data
Now let’s grab some data and frame a question.
Berkeley Admissions
What is the probability of being admitted to UC Berkeley Graduate School?1
I have three variables:
Admitted or not
Gender
The department an applicant applied to.
Admission is the key.
Code
Admit n percent
Admitted 1755 0.3877596
Rejected 2771 0.6122404The proportion is denoted as \hat{p}. We can also do this with table.
A Visual
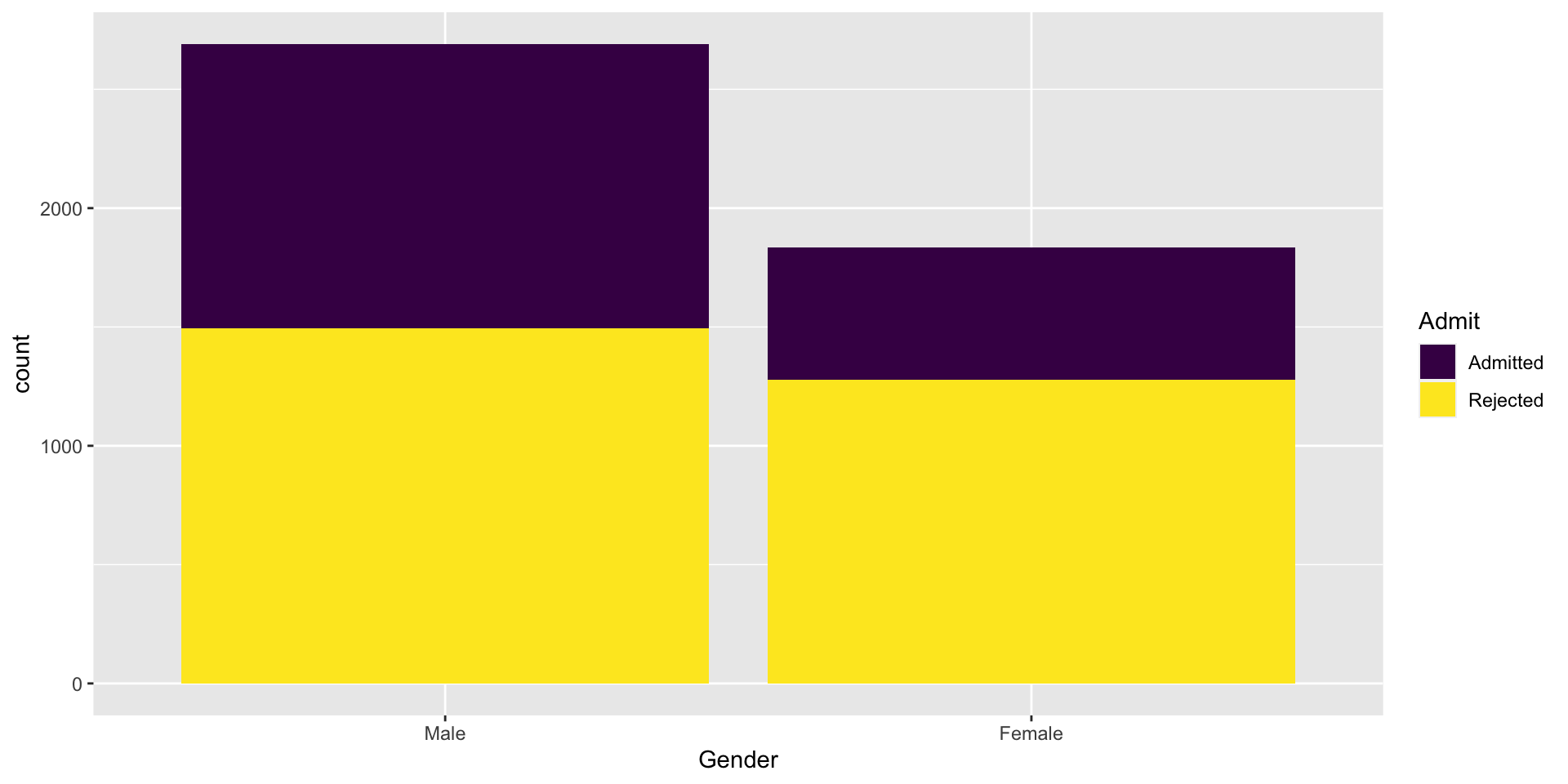
The Idea
Suppose we want to know \pi(Admit) – the probability of being admitted – with 90% probability.
We want to take the data that we saw and use it to infer the likely values of \pi(Admit).
Three interchangeable methods
Resampling/Simulation
Exact binomial computation
A normal approximation
Method 1: Resampling
Suppose I have 4526 chips with 1755 green and 2771 red.
I toss them all on the floor
and pick them up, one at a time,
record the value (green/red),
put the chip back,
[NB: I put it back to avoid getting exactly the same sample every time.]
and repeat 4526 times.
Each count of green chips as a proportion of 4526 total chips constitutes an estimate of the probability of Admit.
I wrote a little program to do just this – ResampleProps.
remotes::install_github("robertwwalker/ResampleProps")Resampling Result
Code
5% 95%
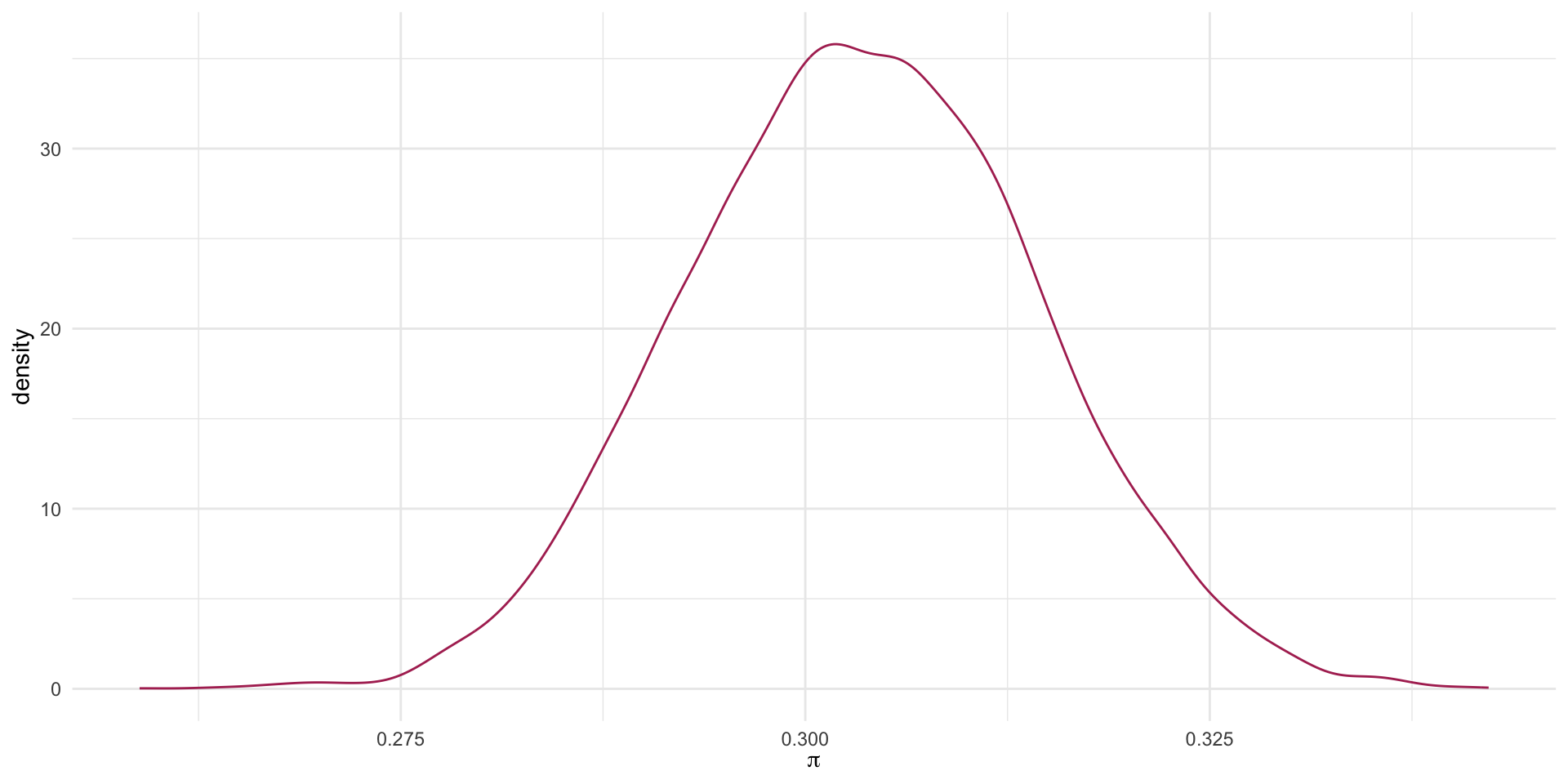
0.3758285 0.3996907 What is our estimate of \pi with 90% confidence?
The probability of admission ranges from 0.376 to 0.4.
A Plot
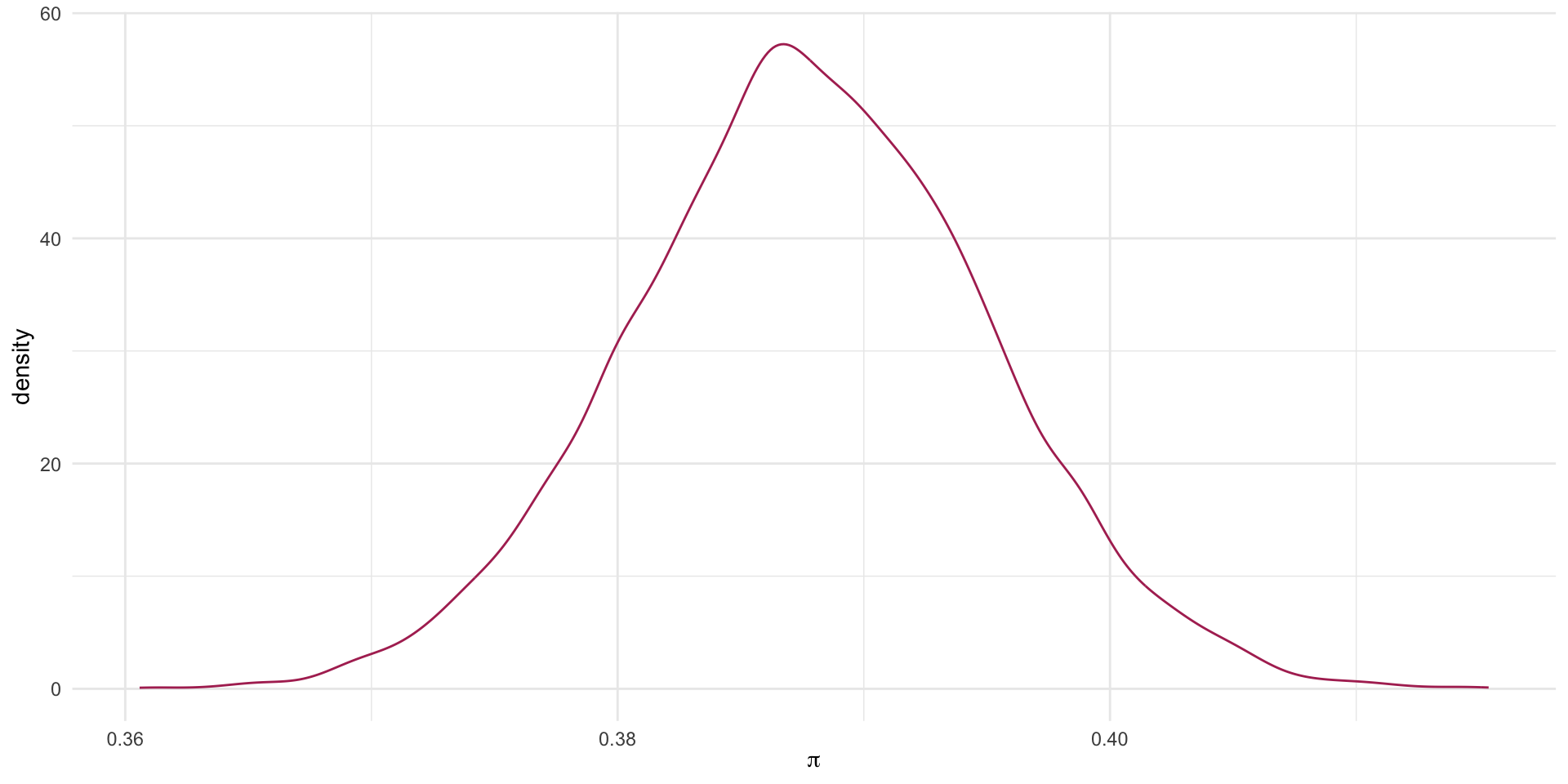
Another Way: Exact binomial
That last procedure is correct but it is overkill.
With probability of 0.05, how small could \pi be to have gotten 1755 of 4526 or more?
With probability 0.95, how big could \pi be to have gotten fewer than 1755 of 4526?
binom.test does exactly this.
binom.test()
Exact binomial test
data: .
number of successes = 1755, number of trials = 4526, p-value < 2.2e-16
alternative hypothesis: true probability of success is not equal to 0.5
90 percent confidence interval:
0.3757924 0.3998340
sample estimates:
probability of success
0.3877596 With 90% probability, now often referred to as 90% confidence to avoid using the word probability twice, the probability of being admitted ranges between 0.3758 and 0.3998.
Illustrating the Binomial
Exact binomial test
data: .
number of successes = 1755, number of trials = 4526, p-value < 2.2e-16
alternative hypothesis: true probability of success is not equal to 0.5
90 percent confidence interval:
0.3757924 0.3998340
sample estimates:
probability of success
0.3877596 [1] 0.05000037 0.95000004Code
Binomial.Search <- data.frame(x=seq(0.33,0.43, by=0.001)) %>% mutate(Too.Low = pbinom(1755, 1755+2771, x), Too.High = 1-pbinom(1754, 1755+2771, x))
Binomial.Search %>% pivot_longer(cols=c(Too.Low,Too.High)) %>% ggplot(., aes(x=x, y=value, color=name)) + geom_line() + geom_hline(aes(yintercept=0.05)) + geom_hline(aes(yintercept=0.95)) + geom_vline(data=Plot.Me, aes(xintercept=.), linetype=3) + labs(title="Using the Binomial to Search", color="Greater/Lesser", x=expression(pi), y="Probability")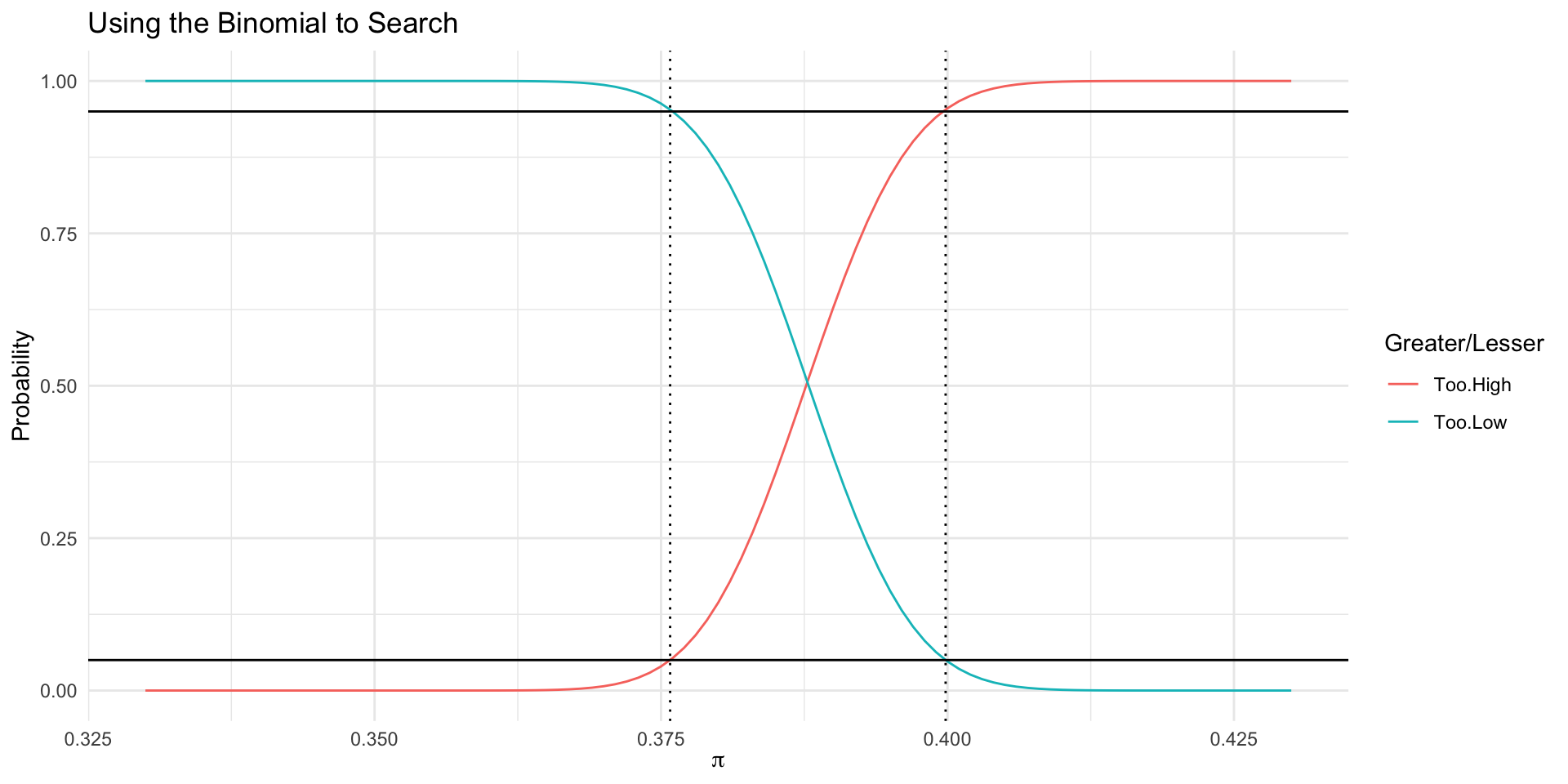
A Third Way: The normal approximation
As long as n and \pi are sufficiently large, we can approximate this with a normal distribution. This will also prove handy for a related reason.
As long as n*\pi > 10, we can write that a standard normal z describes the distribution of \pi, given n – the sample size and \hat{p} – the proportion of yes’s/true’s in the sample.
Pr(\pi) = \hat{p} \pm z \cdot \left( \sqrt{\frac{\hat{p}*(1-\hat{p})}{n}} \right)
R implements this in prop.test. By default, R implements a modified version that corrects for discreteness/continuity. To get the above formula exactly, prop.test(table, correct=FALSE).
A first look at hypotheses
z = \frac{\hat{p} - \pi}{\sqrt{\frac{\pi(1-\pi)}{n}}}
With some proportion calculated from data \hat{p} and some claim about \pi – hereafter called an hypothesis – we can use z to calculate/evaluate the claim. These claims are hypothetical values of \pi. They can be evaluated against a specific alternative considered in three ways:
two-sided, that is \pi = value against not equal.
\pi \geq value against something smaller.
\pi \leq value against something greater.
In the first case, either much bigger or much smaller values could be evidence against equal. The second and third cases are always about smaller and bigger; they are one-sided.
Just as z has metric standard deviations now referred to as standard errors of the proportion, the z here will measure where the data fall with respect to the hypothetical \pi.
With z sufficiently large, and in the proper direction, our hypothetical \pi becomes unlikely given the evidence and should be dismissed.
Three hypothesis tests
The estimate is 15.1 standard errors below the hypothesized value.
Two Sided
\pi = 0.5 against \pi \neq 0.5.
Any |z| > 1.64 – probability 0.05 above and below – is sufficient to dispense with the hypothesis.
One-Sided
\pi \leq 0.5 against \pi \gt 0.5.
With 90% confidence, z > 1.28 is sufficient.
\pi \geq 0.5 against \pi \lt 0.5.
With 90% confidence, z < -1.28 is sufficient.
The normal approximation
1-sample proportions test without continuity correction
data: table(UCBTab$Admit), null probability 0.5
X-squared = 228.07, df = 1, p-value < 2.2e-16
alternative hypothesis: true p is not equal to 0.5
90 percent confidence interval:
0.3759173 0.3997361
sample estimates:
p
0.3877596 R reports the result as z^2 not z which is \chi^2 not normal; we can obtain the z by taking a square root: \sqrt{228.08} \approx 15.1
This approximation yields an estimate of \pi, with 90% confidence, that ranges between 0.376 and 0.4.
A Cool Consequence of Normal
The formal equation defines:
z = \frac{\hat{p} - \pi}{\sqrt{\frac{\pi(1-\pi)}{n}}}
Some language:
Margin of error is \hat{p} - \pi. [MOE]
Confidence: the probability coverage given z [two-sided].
We need a guess at the true \pi because variance/std. dev. depend on \pi. 0.5 is common because it maximizes the variance; we will have enough no matter what the true value.
Algebra allows us to solve for n.
n = \frac{z^2 \cdot \pi(1-\pi)}{(\hat{p} - \pi)^2}
Example
Estimate the probability of supporting an initiative to within 0.03 with 95% confidence. Assume that the probability is 0.5 [it maximizes the variance and renders a conservative estimate – an upper bound on the sample size]
Code
[1] 1068I need 1068 people to estimate support to within plus or minus 0.03 with 95% confidence.
In the Real World

A real poll. They do not have quite enough for a 3% margin of error. But 1006 is enough for a 3.1 percent margin of error…
One More Oddity
What is our estimate of \pi with 90% confidence?
The probability of admission ranges from 0.3758285 to 0.3996907.
If we wish to express this using the normal approximation, a central interval is the observed proportion plus/minus z times the standard error of the proportion – SE(\hat{p}) = \sqrt{\frac{\hat{p}(1-\hat{p})}{n}} NB: The sample value replaces our assumed true value because \pi is to be solved for. For 90%, the probability of admissions ranges from
Back to the Story
Does the probability of admission depend on whether the applicant is Male or Female?
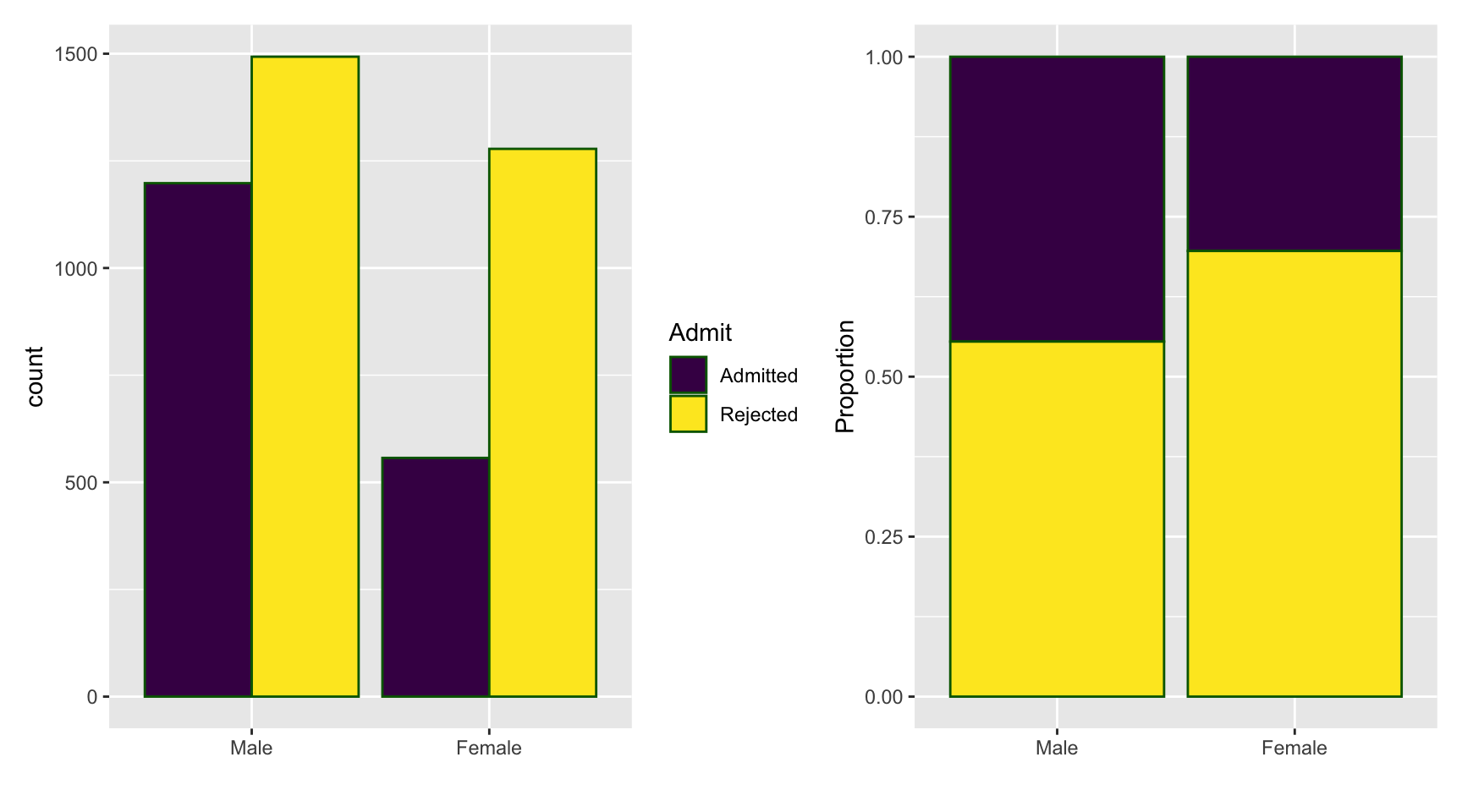
Analysis by Group
Gender Admitted Rejected Total
Male 1198 1493 2691
Female 557 1278 1835 Gender Admitted Rejected
Male 0.4451877 0.5548123
Female 0.3035422 0.6964578Code
UCBTF <- UCBTab %>% filter(Gender=="Female")
UCBTF.Pi <- ResampleProp(UCBTF$Admit, k = 10000) %>% data.frame(Pi.Admit=., Gender=as.character("Female"), frameN=2) # Estimates for females
UCBTM <- UCBTab %>% filter(Gender=="Male")
UCBTM.Pi <- ResampleProp(UCBTM$Admit, k = 10000) %>% data.frame(Pi.Admit=., Gender = as.character("Male"), frameN = 2) # Estimates for malesBinomial Result: Males
Exact binomial test
data: .
number of successes = 1198, number of trials = 2691, p-value =
0.00000001403
alternative hypothesis: true probability of success is not equal to 0.5
90 percent confidence interval:
0.4292930 0.4611706
sample estimates:
probability of success
0.4451877 The probability of being admitted, conditional on being Male, ranges from 0.43 to 0.46 with 90% confidence.
The Thought Experiment: Male
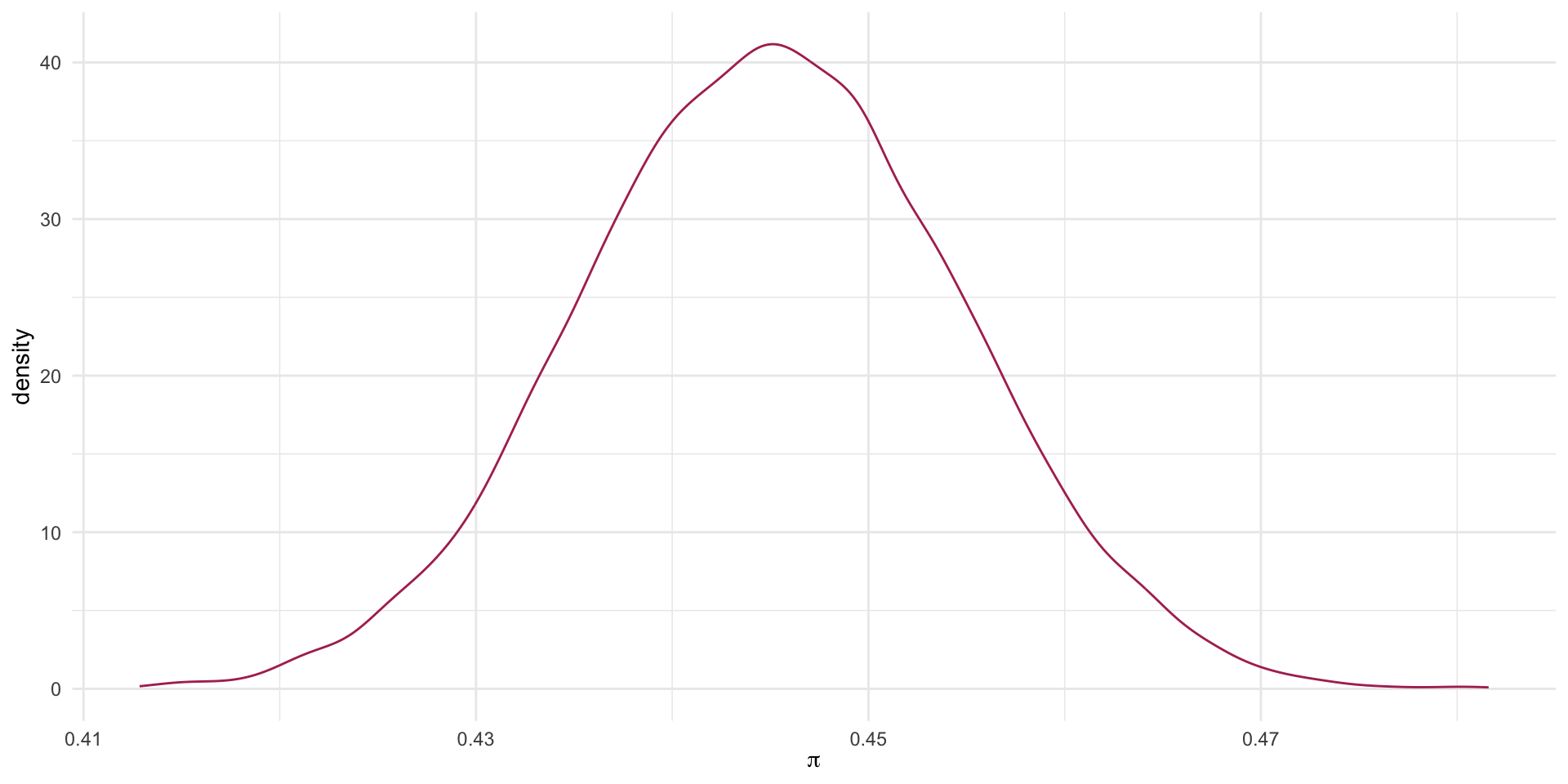
Binomial Result: Females
Exact binomial test
data: .
number of successes = 557, number of trials = 1835, p-value < 2.2e-16
alternative hypothesis: true probability of success is not equal to 0.5
90 percent confidence interval:
0.2858562 0.3216944
sample estimates:
probability of success
0.3035422 The probability of being of Admitted, given a Female, ranges from 0.286 to 0.322 with 90% confidence.
The Thought Experiment: Female
Succinctly
Female: from 0.286 to 0.322
Male: from 0.43 to 0.46
All: from 0.3758 to 0.3998
A Key Visual
Code
UCB.Pi <- bind_rows(UCBTF.Pi, UCBTM.Pi)
UCB.Pi %>% ggplot(., aes(x=Gender, y=Pi.Admit, fill=Gender)) + geom_violin() + geom_label(aes(x=1.5, y=0.375), label="Too small to be male \n Too large to be female?", size=14, fill="black", color="white", inherit.aes = FALSE) + guides(size="none") + labs(x="") + scale_fill_viridis_d()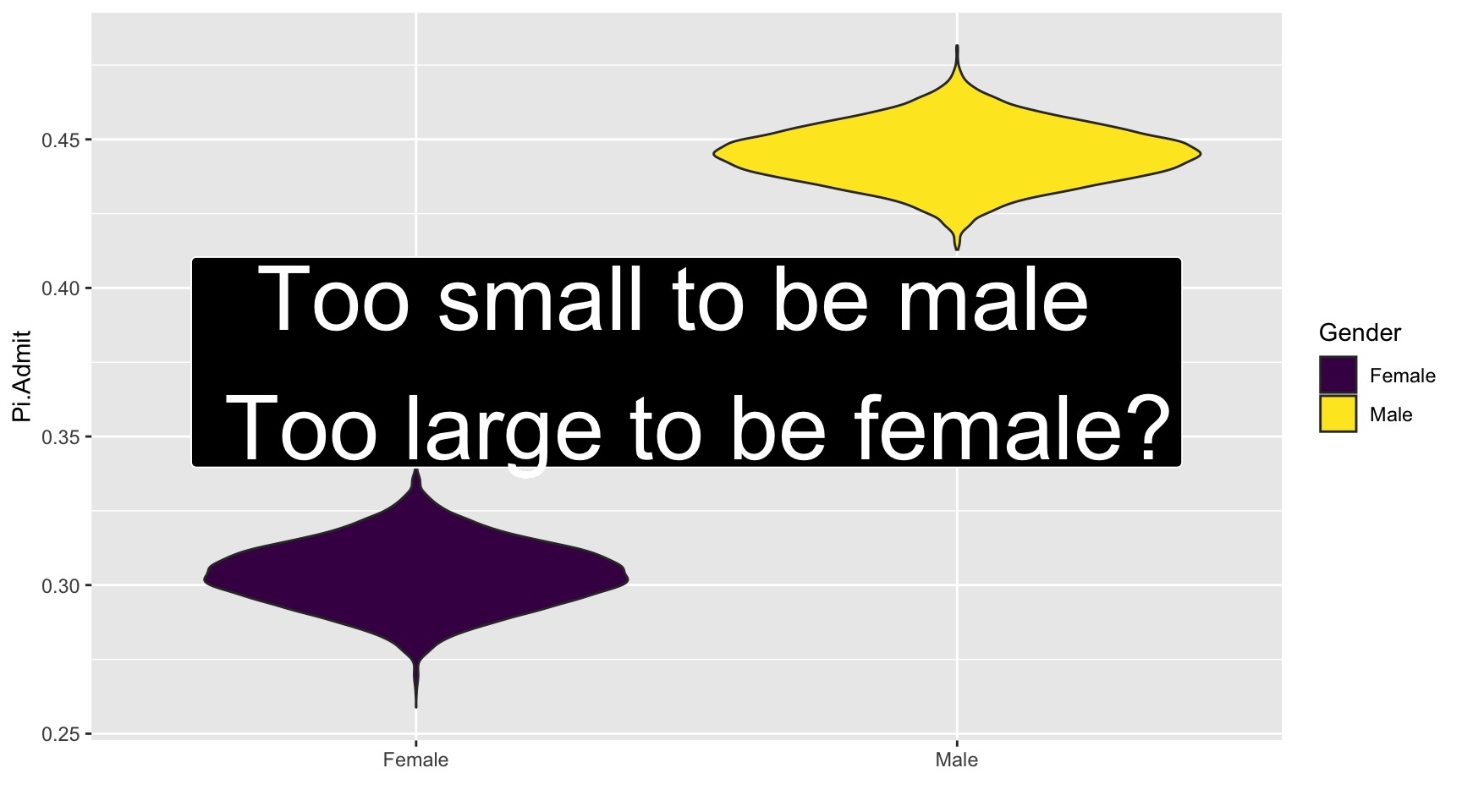
Code
UCB.Pi <- bind_rows(UCBTF.Pi, UCBTM.Pi, RSMP)
UCB.Pi %>% ggplot(., aes(x=Pi.Admit, fill=Gender)) + geom_density(alpha=0.5) + theme_minimal() + scale_fill_viridis_d() + transition_states(frameN, state_length = 40, transition_length = 20) + enter_fade() + exit_fade() -> SaveAnim
anim_save(SaveAnim, renderer = gifski_renderer(height=500, width=1000), filename="img/Anim1.gif")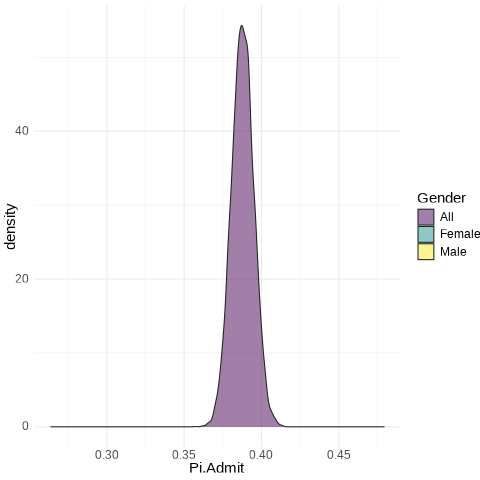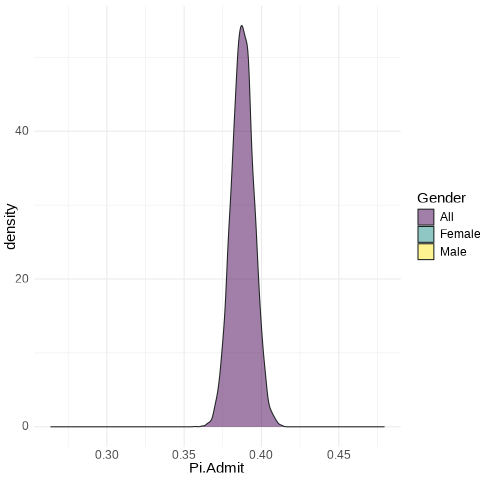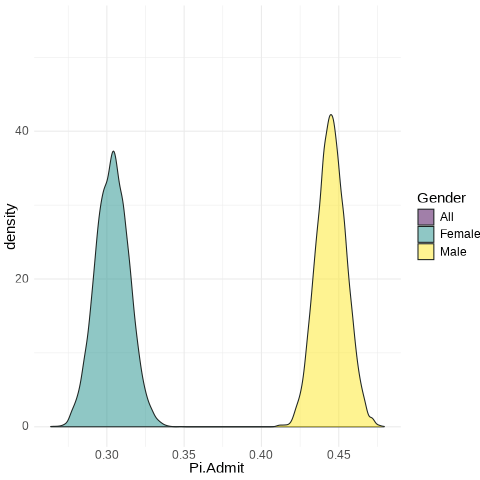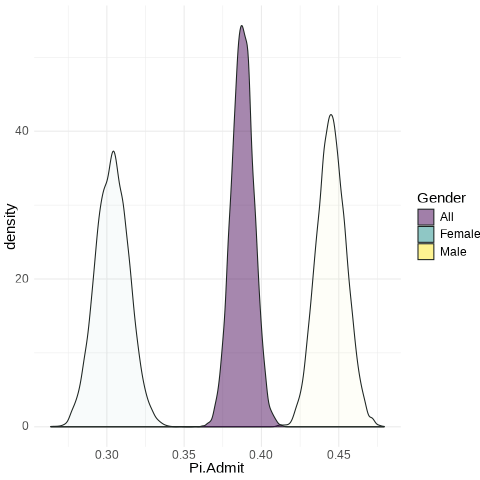
Can We Measure the Difference?
How much more likely are Males to be admitted when compared to Females?
We can take the difference in our estimates for Male and Female.
Code
5% 95%
0.1176662 0.1651037 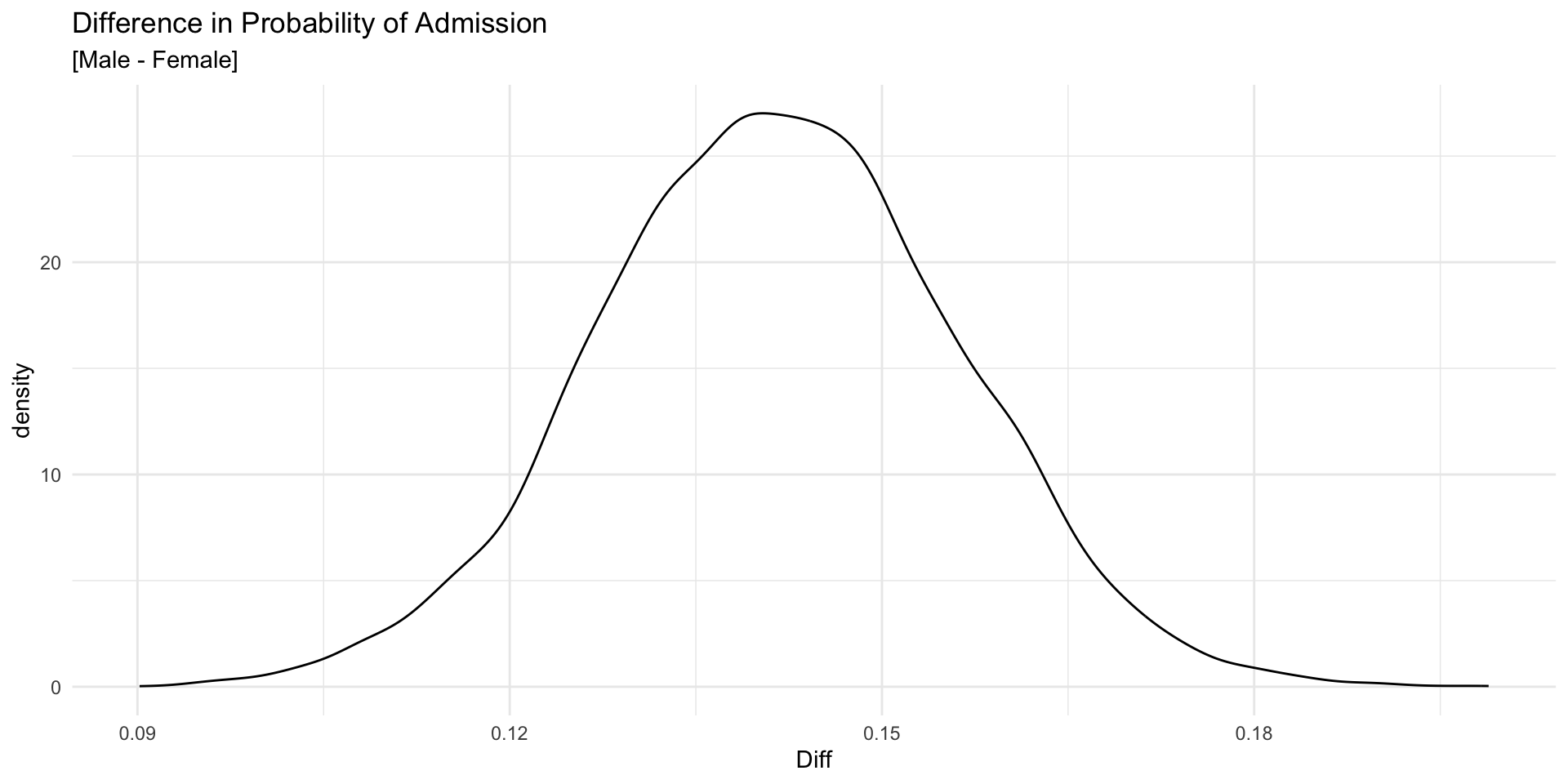
Or Use the Normal Approximation
[\hat{p}_{M} - \hat{p}_{F}] \pm z*\sqrt{ \underbrace{\frac{\hat{p}_{M}(1-\hat{p}_{M})}{n_{M}}}_{Males} + \underbrace{\frac{\hat{p}_{F}(1-\hat{p}_{F})}{n_{F}}}_{Females}}
Code
2-sample test for equality of proportions without continuity correction
data: .
X-squared = 92.205, df = 1, p-value < 2.2e-16
alternative hypothesis: two.sided
90 percent confidence interval:
0.1179805 0.1653103
sample estimates:
prop 1 prop 2
0.4451877 0.3035422 With 90% confidence, a Female is 0.118 to 0.165 [0.1416] less likely to be admitted.
The Standard Error of the Difference
Following something akin to the FOIL method, we can show that the variance of a difference in two random variables is the sum of the variances minus twice the covariance between them.
Var(X - Y) = Var(X) + Var(Y) - 2*Cov(X,Y)
If we assume [or recognize it is generally unknownable] that the male and female samples are independent, the covariance is zero, and the variance of the difference is simply the sum of the variance for male and female, respectively, and zero covariance.
SE(\hat{p}_{M} - \hat{p}_{F}) = \sqrt{ \underbrace{\frac{\hat{p}_{M}(1-\hat{p}_{M})}{n_{M}}}_{Males} + \underbrace{\frac{\hat{p}_{F}(1-\hat{p}_{F})}{n_{F}}}_{Females}}
The chi-square and the test of independence
The sum of k squared standard normal (z or Normal(0,1)) variates has a \chi^2 distribution with k degrees of freedom. Two sample tests of proportions can be reported using the chi-square metric as well. R’s prop.test does this by default.

DADM: Discrete Inference